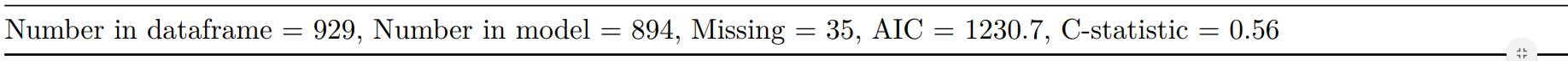The finafit package brings together the day-to-day
functions we use to generate final results tables and plots when
modelling.
I spent many years repeatedly manually copying results from R analyses and built these functions to automate our standard healthcare data workflow. It is particularly useful when undertaking a large study involving multiple different regression analyses. When combined with RMarkdown, the reporting becomes entirely automated. Its design follows Hadley Wickham’s tidy tool manifesto.
Installation and Documentation
Development lives on GitHub.
You can install the finalfit development version from
CRAN with:
install.packages("finalfit")It is recommended that this package is used together with
dplyr, which is a dependent.
Some of the functions require rstan and
boot. These have been left as Suggests rather
than Depends to avoid unnecessary installation. If needed,
they can be installed in the normal way:
install.packages("rstan")
install.packages("boot")To
install
off-line (or in a Safe Haven), download the zip file and use
devtools::install_local().
Main Features
1. Summarise variables/factors by a categorical variable
summary_factorlist() is a wrapper used to aggregate any
number of explanatory variables by a single variable of
interest. This is often “Table 1” of a published study. When
categorical, the variable of interest can have a maximum of five levels.
It uses Hmisc::summary.formula().
library(finalfit)
library(dplyr)
# Load example dataset, modified version of survival::colon
data(colon_s)
# Table 1 - Patient demographics by variable of interest ----
explanatory = c("age", "age.factor", "sex.factor", "obstruct.factor")
dependent = "perfor.factor" # Bowel perforation
colon_s %>%
summary_factorlist(dependent, explanatory,
p=TRUE, add_dependent_label=TRUE) -> t1
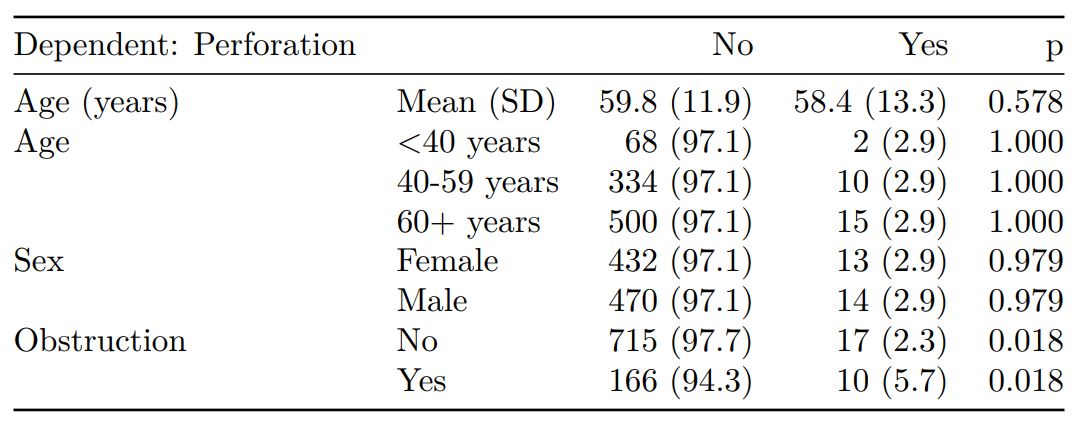
knitr::kable(t1, row.names=FALSE, align=c("l", "l", "r", "r", "r"))| Dependent: Perforation | No | Yes | p | |
|---|---|---|---|---|
| Age (years) | Mean (SD) | 59.8 (11.9) | 58.4 (13.3) | 0.542 |
| Age | <40 years | 68 (7.5) | 2 (7.4) | 1.000 |
| 40-59 years | 334 (37.0) | 10 (37.0) | ||
| 60+ years | 500 (55.4) | 15 (55.6) | ||
| Sex | Female | 432 (47.9) | 13 (48.1) | 1.000 |
| Male | 470 (52.1) | 14 (51.9) | ||
| Obstruction | No | 715 (81.2) | 17 (63.0) | 0.035 |
| Yes | 166 (18.8) | 10 (37.0) |
When exported to PDF:
See other options relating to inclusion of missing data, mean vs. median for continuous variables, column vs. row proportions, include a total column etc.
summary_factorlist() is also commonly used to summarise
any number of variables by an outcome variable (say
dead yes/no).
# Table 2 - 5 yr mortality ----
explanatory = c("age.factor", "sex.factor", "obstruct.factor")
dependent = 'mort_5yr'
colon_s %>%
summary_factorlist(dependent, explanatory,
p=TRUE, add_dependent_label=TRUE) -> t2
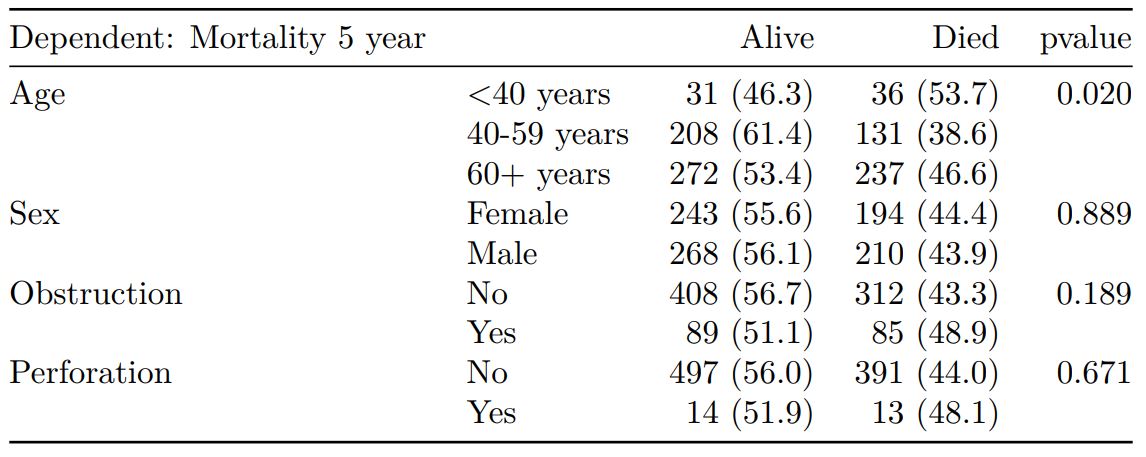
knitr::kable(t2, row.names=FALSE, align=c("l", "l", "r", "r", "r"))| Dependent: Mortality 5 year | Alive | Died | p | |
|---|---|---|---|---|
| Age | <40 years | 31 (6.1) | 36 (8.9) | 0.020 |
| 40-59 years | 208 (40.7) | 131 (32.4) | ||
| 60+ years | 272 (53.2) | 237 (58.7) | ||
| Sex | Female | 243 (47.6) | 194 (48.0) | 0.941 |
| Male | 268 (52.4) | 210 (52.0) | ||
| Obstruction | No | 408 (82.1) | 312 (78.6) | 0.219 |
| Yes | 89 (17.9) | 85 (21.4) |
Tables can be knitted to PDF, Word or html documents. We do this in RStudio from a .Rmd document.
2. Summarise regression model results in final table format
The second main feature is the ability to create final tables for
linear lm(), logistic glm(), hierarchical
logistic lme4::glmer() and Cox proportional hazards
survival::coxph() regression models.
The finalfit() “all-in-one” function takes a single
dependent variable with a vector of explanatory variable names
(continuous or categorical variables) to produce a final table for
publication including summary statistics, univariable and multivariable
regression analyses. The first columns are those produced by
summary_factorist(). The appropriate regression model is
chosen on the basis of the dependent variable type and other arguments
passed.
Logistic regression: glm()
Of the form:
glm(depdendent ~ explanatory, family="binomial")
explanatory = c("age.factor", "sex.factor", "obstruct.factor", "perfor.factor")
dependent = 'mort_5yr'
colon_s %>%
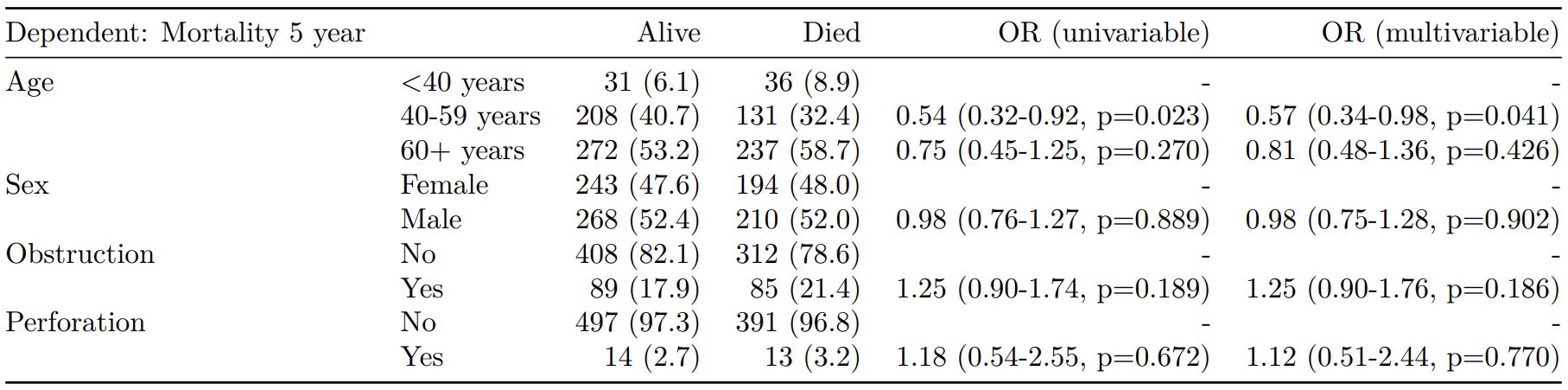
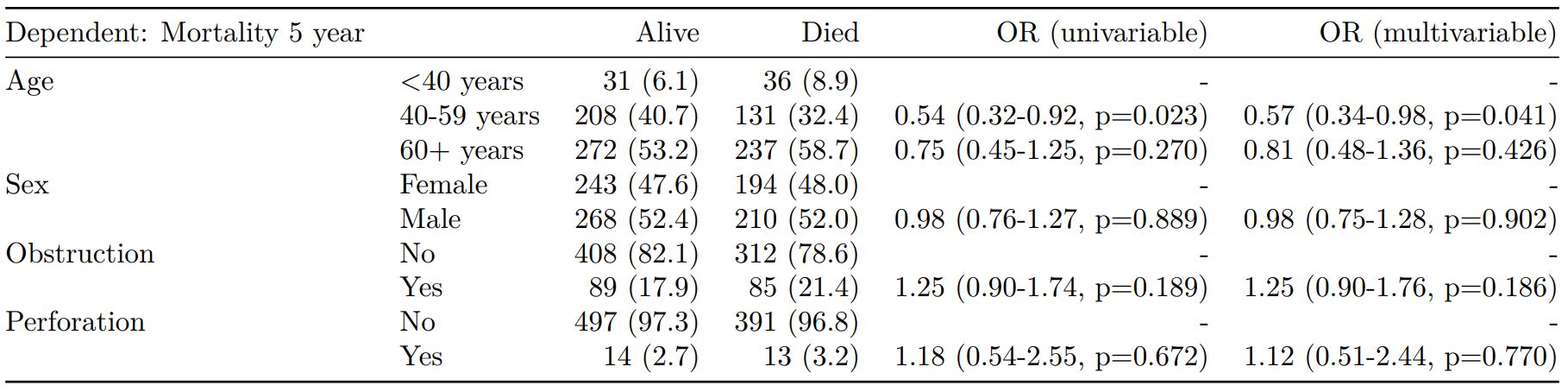
finalfit(dependent, explanatory) -> t3
knitr::kable(t3, row.names=FALSE, align=c("l", "l", "r", "r", "r", "r"))| Dependent: Mortality 5 year | Alive | Died | OR (univariable) | OR (multivariable) | |
|---|---|---|---|---|---|
| Age | <40 years | 31 (46.3) | 36 (53.7) | - | - |
| 40-59 years | 208 (61.4) | 131 (38.6) | 0.54 (0.32-0.92, p=0.023) | 0.57 (0.34-0.98, p=0.041) | |
| 60+ years | 272 (53.4) | 237 (46.6) | 0.75 (0.45-1.25, p=0.270) | 0.81 (0.48-1.36, p=0.426) | |
| Sex | Female | 243 (55.6) | 194 (44.4) | - | - |
| Male | 268 (56.1) | 210 (43.9) | 0.98 (0.76-1.27, p=0.889) | 0.98 (0.75-1.28, p=0.902) | |
| Obstruction | No | 408 (56.7) | 312 (43.3) | - | - |
| Yes | 89 (51.1) | 85 (48.9) | 1.25 (0.90-1.74, p=0.189) | 1.25 (0.90-1.76, p=0.186) | |
| Perforation | No | 497 (56.0) | 391 (44.0) | - | - |
| Yes | 14 (51.9) | 13 (48.1) | 1.18 (0.54-2.55, p=0.672) | 1.12 (0.51-2.44, p=0.770) |
Logistic regression with reduced model: glm()
Where a multivariable model contains a subset of the variables included specified in the full univariable set, this can be specified.
explanatory = c("age.factor", "sex.factor", "obstruct.factor", "perfor.factor")
explanatory_multi = c("age.factor", "obstruct.factor")
dependent = 'mort_5yr'
colon_s %>%
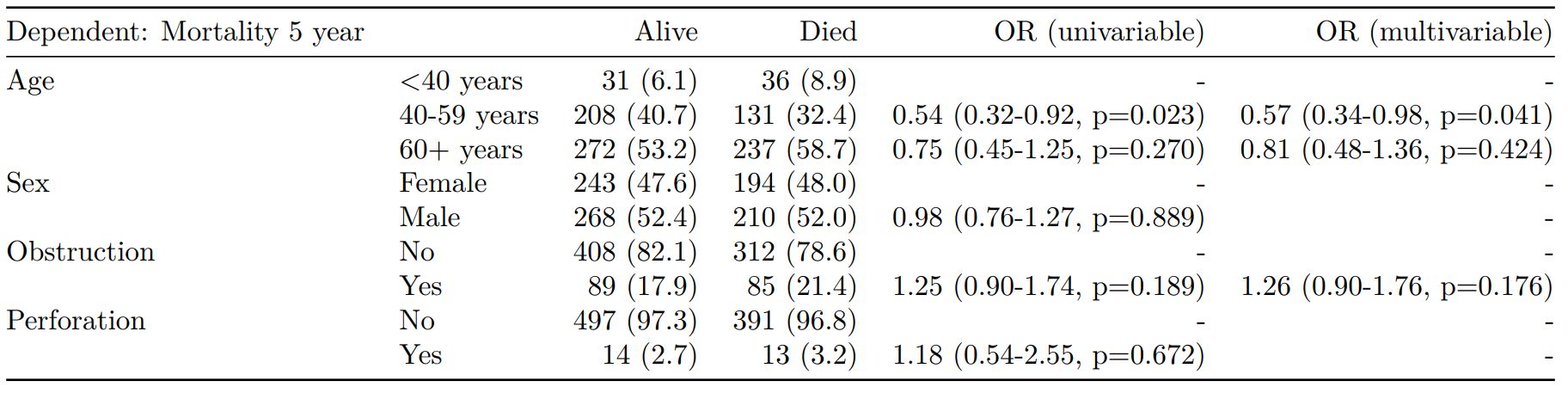
finalfit(dependent, explanatory, explanatory_multi) -> t4
knitr::kable(t4, row.names=FALSE, align=c("l", "l", "r", "r", "r", "r"))| Dependent: Mortality 5 year | Alive | Died | OR (univariable) | OR (multivariable) | |
|---|---|---|---|---|---|
| Age | <40 years | 31 (46.3) | 36 (53.7) | - | - |
| 40-59 years | 208 (61.4) | 131 (38.6) | 0.54 (0.32-0.92, p=0.023) | 0.57 (0.34-0.98, p=0.041) | |
| 60+ years | 272 (53.4) | 237 (46.6) | 0.75 (0.45-1.25, p=0.270) | 0.81 (0.48-1.36, p=0.424) | |
| Sex | Female | 243 (55.6) | 194 (44.4) | - | - |
| Male | 268 (56.1) | 210 (43.9) | 0.98 (0.76-1.27, p=0.889) | - | |
| Obstruction | No | 408 (56.7) | 312 (43.3) | - | - |
| Yes | 89 (51.1) | 85 (48.9) | 1.25 (0.90-1.74, p=0.189) | 1.26 (0.90-1.76, p=0.176) | |
| Perforation | No | 497 (56.0) | 391 (44.0) | - | - |
| Yes | 14 (51.9) | 13 (48.1) | 1.18 (0.54-2.55, p=0.672) | - |
Mixed effects logistic regression: lme4::glmer()
Of the form:
lme4::glmer(dependent ~ explanatory + (1 | random_effect), family="binomial")
Hierarchical/mixed effects/multilevel logistic regression models can
be specified using the argument random_effect. At the
moment it is just set up for random intercepts
(i.e. (1 | random_effect), but in the future I’ll adjust
this to accommodate random gradients if needed
(i.e. (variable1 | variable2).
explanatory = c("age.factor", "sex.factor", "obstruct.factor", "perfor.factor")
explanatory_multi = c("age.factor", "obstruct.factor")
random_effect = "hospital"
dependent = 'mort_5yr'
colon_s %>%
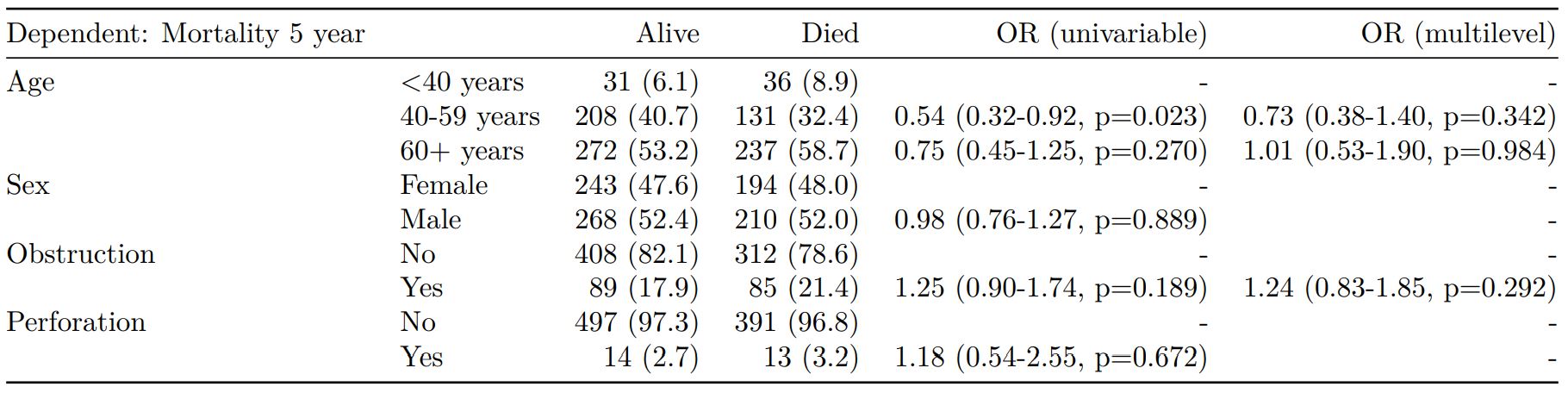
finalfit(dependent, explanatory, explanatory_multi, random_effect) -> t5
knitr::kable(t5, row.names=FALSE, align=c("l", "l", "r", "r", "r", "r"))| Dependent: Mortality 5 year | Alive | Died | OR (univariable) | OR (multilevel) | |
|---|---|---|---|---|---|
| Age | <40 years | 31 (46.3) | 36 (53.7) | - | - |
| 40-59 years | 208 (61.4) | 131 (38.6) | 0.54 (0.32-0.92, p=0.023) | 0.73 (0.38-1.40, p=0.342) | |
| 60+ years | 272 (53.4) | 237 (46.6) | 0.75 (0.45-1.25, p=0.270) | 1.01 (0.53-1.90, p=0.984) | |
| Sex | Female | 243 (55.6) | 194 (44.4) | - | - |
| Male | 268 (56.1) | 210 (43.9) | 0.98 (0.76-1.27, p=0.889) | - | |
| Obstruction | No | 408 (56.7) | 312 (43.3) | - | - |
| Yes | 89 (51.1) | 85 (48.9) | 1.25 (0.90-1.74, p=0.189) | 1.24 (0.83-1.85, p=0.292) | |
| Perforation | No | 497 (56.0) | 391 (44.0) | - | - |
| Yes | 14 (51.9) | 13 (48.1) | 1.18 (0.54-2.55, p=0.672) | - |
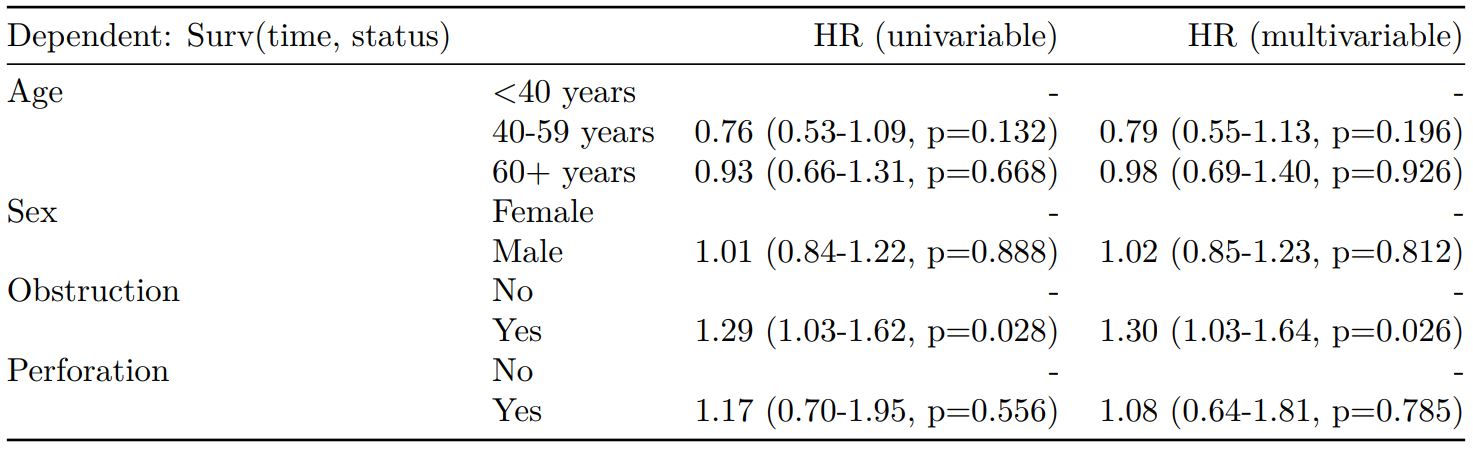
Cox proportional hazards: survival::coxph()
Of the form:
survival::coxph(dependent ~ explanatory)
explanatory = c("age.factor", "sex.factor", "obstruct.factor", "perfor.factor")
dependent = "Surv(time, status)"
colon_s %>%
finalfit(dependent, explanatory) -> t6
knitr::kable(t6, row.names=FALSE, align=c("l", "l", "r", "r", "r", "r"))| Dependent: Surv(time, status) | all | HR (univariable) | HR (multivariable) | |
|---|---|---|---|---|
| Age | <40 years | 70 (7.5) | - | - |
| 40-59 years | 344 (37.0) | 0.76 (0.53-1.09, p=0.132) | 0.79 (0.55-1.13, p=0.196) | |
| 60+ years | 515 (55.4) | 0.93 (0.66-1.31, p=0.668) | 0.98 (0.69-1.40, p=0.926) | |
| Sex | Female | 445 (47.9) | - | - |
| Male | 484 (52.1) | 1.01 (0.84-1.22, p=0.888) | 1.02 (0.85-1.23, p=0.812) | |
| Obstruction | No | 732 (80.6) | - | - |
| Yes | 176 (19.4) | 1.29 (1.03-1.62, p=0.028) | 1.30 (1.03-1.64, p=0.026) | |
| Perforation | No | 902 (97.1) | - | - |
| Yes | 27 (2.9) | 1.17 (0.70-1.95, p=0.556) | 1.08 (0.64-1.81, p=0.785) |
Add common model metrics to output
metrics=TRUE provides common model metrics. The output
is a list of two dataframes. Note chunk specification for output
below.
explanatory = c("age.factor", "sex.factor",
"obstruct.factor", "perfor.factor")
dependent = 'mort_5yr'
colon_s %>%
finalfit(dependent, explanatory, metrics=TRUE) -> t7
knitr::kable(t7[[1]], row.names=FALSE, align=c("l", "l", "r", "r", "r", "r"))| Dependent: Mortality 5 year | Alive | Died | OR (univariable) | OR (multivariable) | |
|---|---|---|---|---|---|
| Age | <40 years | 31 (46.3) | 36 (53.7) | - | - |
| 40-59 years | 208 (61.4) | 131 (38.6) | 0.54 (0.32-0.92, p=0.023) | 0.57 (0.34-0.98, p=0.041) | |
| 60+ years | 272 (53.4) | 237 (46.6) | 0.75 (0.45-1.25, p=0.270) | 0.81 (0.48-1.36, p=0.426) | |
| Sex | Female | 243 (55.6) | 194 (44.4) | - | - |
| Male | 268 (56.1) | 210 (43.9) | 0.98 (0.76-1.27, p=0.889) | 0.98 (0.75-1.28, p=0.902) | |
| Obstruction | No | 408 (56.7) | 312 (43.3) | - | - |
| Yes | 89 (51.1) | 85 (48.9) | 1.25 (0.90-1.74, p=0.189) | 1.25 (0.90-1.76, p=0.186) | |
| Perforation | No | 497 (56.0) | 391 (44.0) | - | - |
| Yes | 14 (51.9) | 13 (48.1) | 1.18 (0.54-2.55, p=0.672) | 1.12 (0.51-2.44, p=0.770) |
knitr::kable(t7[[2]], row.names=FALSE, col.names="")| Number in dataframe = 929, Number in model = 894, Missing = 35, AIC = 1230.7, C-statistic = 0.56, H&L = Chi-sq(8) 5.69 (p=0.682) |
Combine multiple models into single table
Rather than going all-in-one, any number of subset models can be
manually added on to a summary_factorlist() table using
finalfit_merge(). This is particularly useful when models
take a long-time to run or are complicated.
Note the requirement for fit_id=TRUE in
summary_factorlist(). fit2df extracts,
condenses, and add metrics to supported models.
explanatory = c("age.factor", "sex.factor", "obstruct.factor", "perfor.factor")
explanatory_multi = c("age.factor", "obstruct.factor")
random_effect = "hospital"
dependent = 'mort_5yr'
# Separate tables
colon_s %>%
summary_factorlist(dependent,
explanatory, fit_id=TRUE) -> example.summary
colon_s %>%
glmuni(dependent, explanatory) %>%
fit2df(estimate_suffix=" (univariable)") -> example.univariable
colon_s %>%
glmmulti(dependent, explanatory) %>%
fit2df(estimate_suffix=" (multivariable)") -> example.multivariable
colon_s %>%
glmmixed(dependent, explanatory, random_effect) %>%
fit2df(estimate_suffix=" (multilevel)") -> example.multilevel
# Pipe together
example.summary %>%
finalfit_merge(example.univariable) %>%
finalfit_merge(example.multivariable) %>%
finalfit_merge(example.multilevel, last_merge = TRUE) %>%
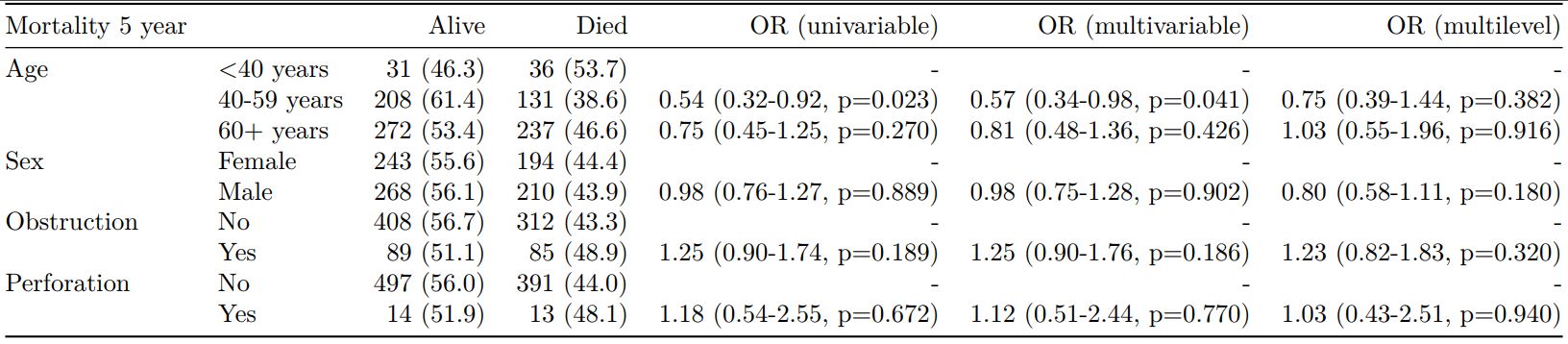
dependent_label(colon_s, dependent, prefix="") -> t8 # place dependent variable label
knitr::kable(t8, row.names=FALSE, align=c("l", "l", "r", "r", "r", "r", "r"))| Mortality 5 year | Alive | Died | OR (univariable) | OR (multivariable) | OR (multilevel) | |
|---|---|---|---|---|---|---|
| Age | <40 years | 31 (6.1) | 36 (8.9) | - | - | - |
| 40-59 years | 208 (40.7) | 131 (32.4) | 0.54 (0.32-0.92, p=0.023) | 0.57 (0.34-0.98, p=0.041) | 0.75 (0.39-1.44, p=0.382) | |
| 60+ years | 272 (53.2) | 237 (58.7) | 0.75 (0.45-1.25, p=0.270) | 0.81 (0.48-1.36, p=0.426) | 1.03 (0.55-1.96, p=0.916) | |
| Sex | Female | 243 (47.6) | 194 (48.0) | - | - | - |
| Male | 268 (52.4) | 210 (52.0) | 0.98 (0.76-1.27, p=0.889) | 0.98 (0.75-1.28, p=0.902) | 0.80 (0.58-1.11, p=0.180) | |
| Obstruction | No | 408 (82.1) | 312 (78.6) | - | - | - |
| Yes | 89 (17.9) | 85 (21.4) | 1.25 (0.90-1.74, p=0.189) | 1.25 (0.90-1.76, p=0.186) | 1.23 (0.82-1.83, p=0.320) | |
| Perforation | No | 497 (97.3) | 391 (96.8) | - | - | - |
| Yes | 14 (2.7) | 13 (3.2) | 1.18 (0.54-2.55, p=0.672) | 1.12 (0.51-2.44, p=0.770) | 1.03 (0.43-2.51, p=0.940) |
Bayesian logistic regression: with stan
Our own particular rstan models are supported and will
be documented in the future. Broadly, if you are running (hierarchical)
logistic regression models in Stan with
coefficients specified as a vector labelled beta, then
fit2df() will work directly on the stanfit
object in a similar manner to if it was a glm or
glmerMod object.
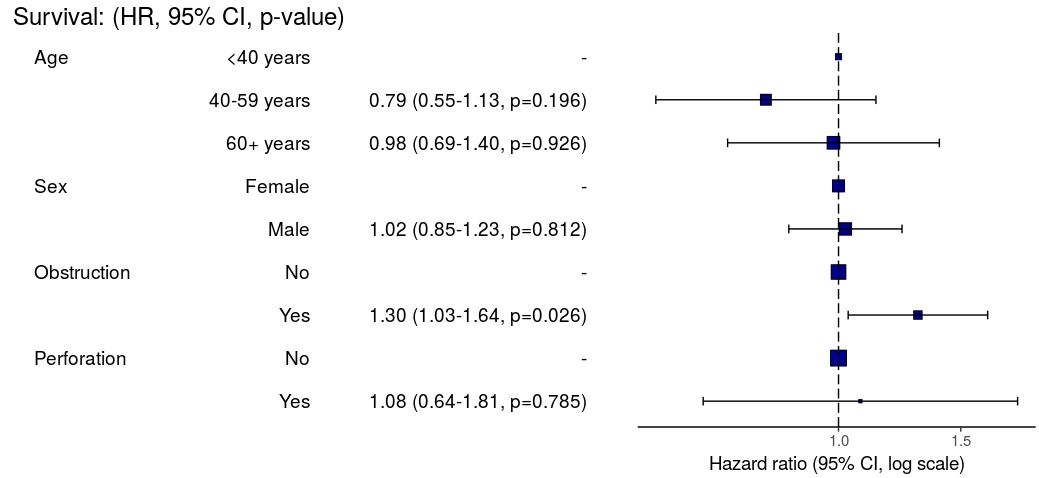
3. Summarise regression model results in plot
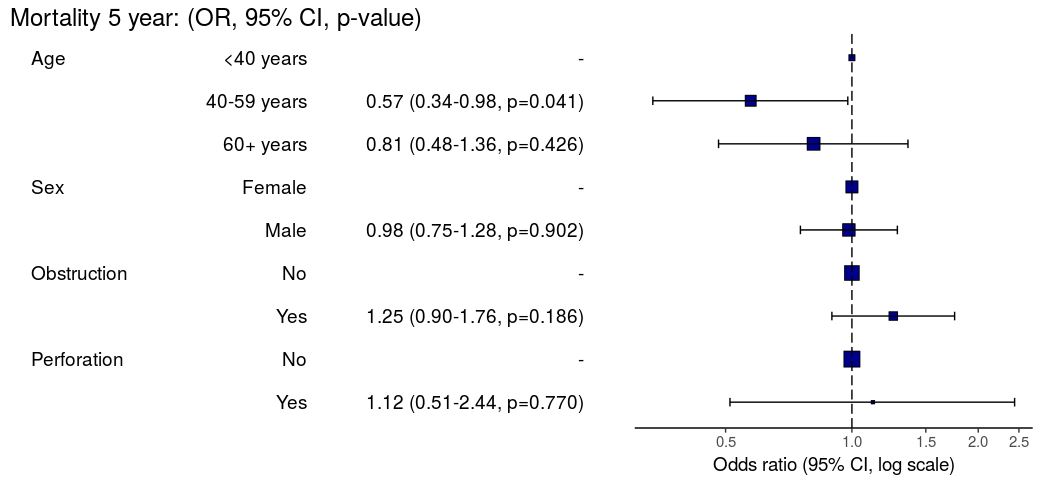
Models can be summarized with odds ratio/hazard ratio plots using
or_plot, hr_plot and
surv_plot.
OR plot
HR plot
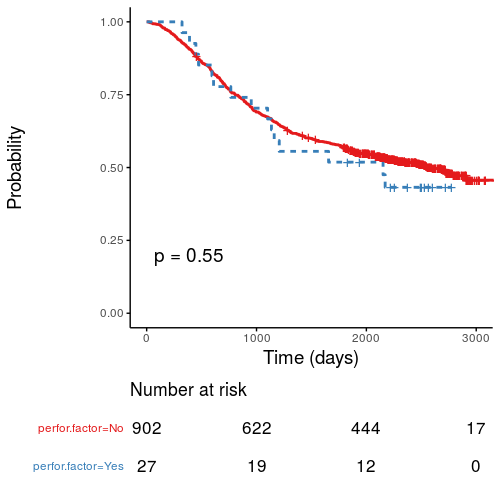
Kaplan-Meier survival plots
KM plots can be produced using the
library(survminer)
Notes
Use ff_label() to assign labels to variables for tables
and plots.
Export dataframe tables directly or to
R Markdown
knitr::kable().
Note wrapper missing_pattern() is also useful. Wraps
mice::md.pattern.
colon_s %>%
missing_pattern(dependent, explanatory)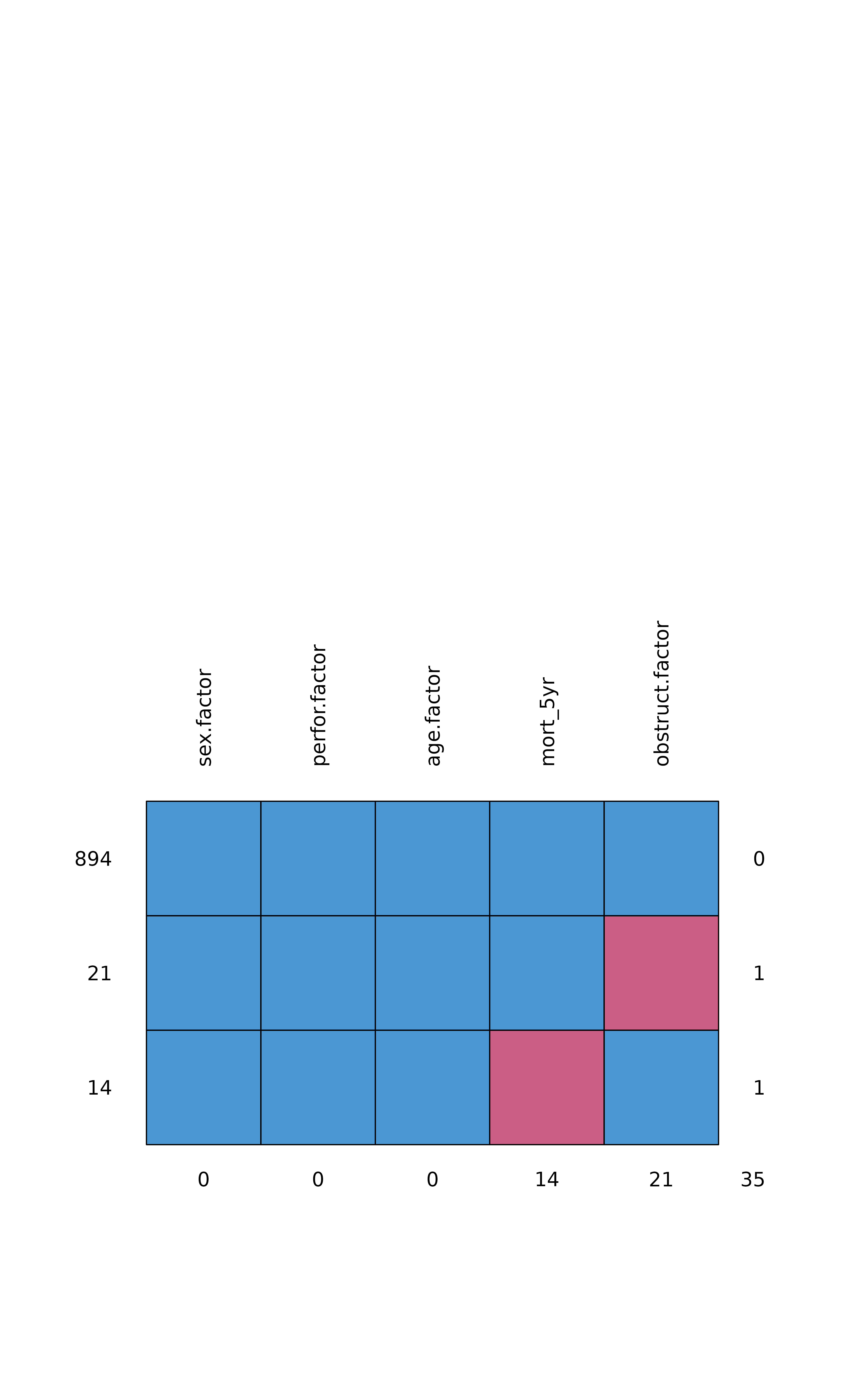
#> sex.factor perfor.factor age.factor mort_5yr obstruct.factor
#> 894 1 1 1 1 1 0
#> 21 1 1 1 1 0 1
#> 14 1 1 1 0 1 1
#> 0 0 0 14 21 35Development will be on-going, but any input appreciated.